|
geom_smooth( aes( color = "y ~ log(x)", fill = "y ~ log(x)"), method = "lm", formula = y ~ log(x), se = FALSE, # Plot the line only (without confidence bands) fullrange = TRUE # The fit spans the full range of the horizontal axis ) + geom_point( aes( color = region, fill = region), size = 2.5, alpha = 0.5, shape = 21 # This is a dot with both border (color) and fill. # F-statistic: 6.961 on 2 and 27 DF, p-value: 0.# Okabe Ito colors # The last color is used for the regression fit. # Residual standard error: 1.999 on 27 degrees of freedom Use summary() to extract estimated cowfficients and standard errors. The results of the model fit are saved in an lm() object productivit圜urve. Wrap the squared term with I() so that the arithmetic ( productivity^2) is done before it is entered as a term in the model. Use lm() to fit a quadratic function to the data. Many of the functions that can be used to extract results from a saved lm object work in the same way when applied to an nls object, such as predict, residuals, and coef. # Achieved convergence tolerance: 6.672e-06 # Residual standard error: 0.1133 on 11 degrees of freedom The first function below carried out the model fit and save the results in an R object named phytoCurve. The value of the initial guess is not so important – here we choose a=1 and b=1 as initial guesses. The function includes an argument providing an initial guess of parameter values. We use “a” and “b” to indicate the parameters of the Michaelis-Menten curve to estimate. Provide a formula that also includes symbols (letters) for the parameters to be estimated. Use the nls (nonlinear least squares) to fit the curve. Labs(x = "Midparent color score", y = "Residuals") + Geom_hline(yintercept=0, color = "black") + Geom_point(size = 3, col = "firebrick") + Ggplot(capColor, aes(midparentColorScore, residuals)) + capColorRegression <- lm(offspringColorScore ~ midparentColorScore, data = capColor)ĬapColor$residuals <- residuals(capColorRegression) Then plot the residuals against the explanatory variable using ggplot(). First, fit the linear regression model using lm() and save the residuals in the data frame. 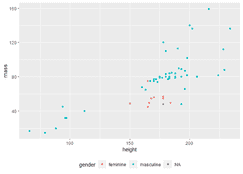
Our Figure 17.5-4 can be reproduced using the following commands. capColorRegression <- lm(offspringColorScore ~ midparentColorScore, data = capColor) Head(capColor) # midparentColorScore offspringColorScoreĪ quick residual plot can be obtained by plotting an lm() object with the ``plot()``` functon. 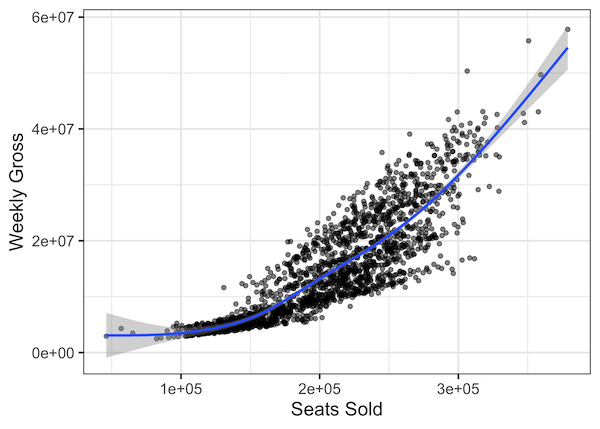
capColor <- read.csv(url(""), stringsAsFactors = FALSE) The plot on the left is from a data set of blue tit cap color of offspring and parents. # Residual standard error: 1.669 on 30 degrees of freedom Save the results into a model object, and then use other commands to extract the quantities wanted. Fit the linear regression to the data using least squares.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
